Content
Research overview

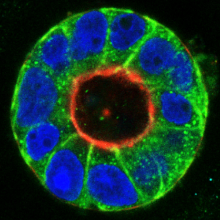
Tumor cells are characterized by complex molecular changes that lead to aberrant survival, growth and motility. Our team investigates the interplay of tumor suppressive and oncogenic signaling networks in cellular transformation. A particular focus is on the signaling pathways that contribute to cancer cell metastasis. Using 3D tissue culture models, we study the molecular interactions with the tumor cell environment, aiming to identify new therapeutic points of intervention. We further employ advanced genetic engineering approaches, primary (co)culture models in combination with single cell and state-of-the-art imaging techniques.
Selected Current Projects
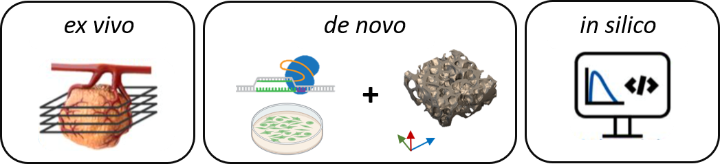
The 3R-US network develops a versatile tissue culture platform for drug testing to replace, reduce and refine (3R) animal experiments. Currently, most studies evaluating anti-cancer therapeutics proceed from human cell culture to preclinical cell line or patient derived mouse tumor models (CDX, PDX). These CDX/PDX models poorly recapitulate the complexity, heterogeneity, and physiology of human tumors. With interdisciplinary partners in cell biology, biomedical engineering and oncology, 3R-US will therefore advance methods that enable the long-term ex vivo cultivation of mouse and human tumor tissue slices. New (targeted) therapeutics and combination therapies will be evaluated in these slice cultures and the mechanisms of drug action investigated. Furthermore, 3R-US will develop de novo human tumor models using 3D printing to study heterogeneous cell interactions and drug responses. Finally, 3R-US will use the data derived from the ex vivo and de novo approaches to establish and validate in silico tumor models that reliably predict the pharmacokinetics and therapy response to anti-cancer drugs.
https://www.verbund.uni-stuttgart.de/3r-biomedicus/3r-research/3r-us/
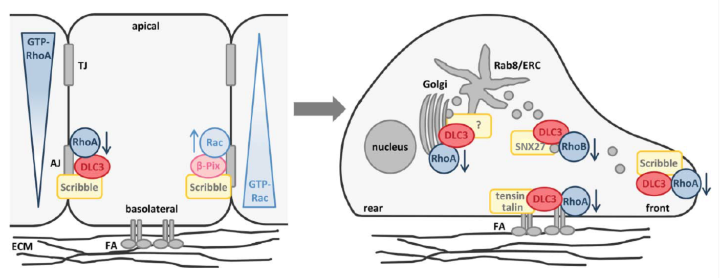
The Rho GTPase activating (GAP) protein Deleted in Liver Cancer 1 (DLC1) has emerged as an important tumor suppressor, whose downregulation in various types of cancer may be as common as that of p53. Our work has shown that DLC1 loss facilitates the aberrant migration of breast cancer cells, whereas expression of the family member DLC3 is crucial for the establishment and maintenance of cell polarity and cell-cell adhesions. While Rho signaling is regulated negatively by GAP proteins on the one hand, it is regulated positively by GEF proteins on the other. We are especially interested in unraveling how specific GEF-GAP networks regulate spatiotemporal Rho signaling and how dysregulation contributes to the metastatic behavior of cancer cells. As partner within the H2020 SECRET Innovative Training Network, we aim to identify drug vulnerabilities of cancer cells with DLC downregulation.
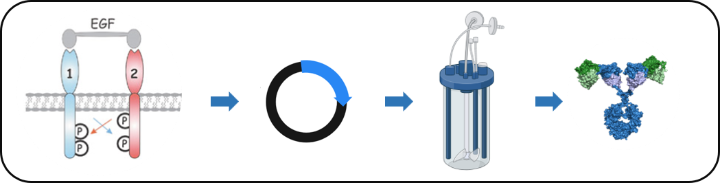
The ErbB family of receptor tyrosine kinases (RTKs) plays a well-established role in cancer development and progression. ErbB2/HER2, for example, is amplified and/or overexpressed in approximately 25% of breast cancer patients, correlating with poor clinical prognosis, whereas EGFR/ErbB1 expression is elevated in 70% of triple-negative breast cancers. Although these receptors are prime targets for pharmacological intervention in the clinic, cancer cells are often non-responsive or become resistant to treatment. In collaboration with the lab of Prof. Roland Kontermann we are developing novel RTK-based targeted approaches for personalized treatment of solid cancers.
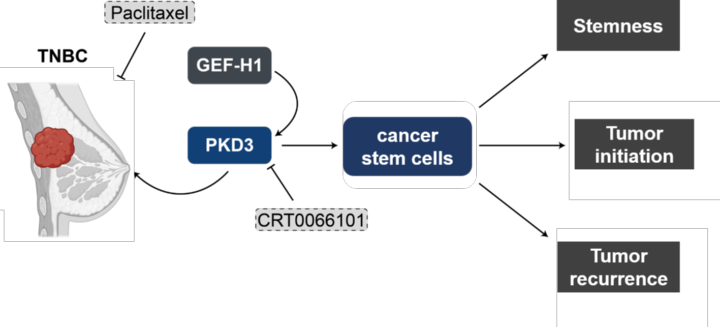
Triple-negative breast cancer (TNBC) is an especially aggressive form of breast cancer for which targeted therapies are still in their infancy. We identified the serine-threonine kinase PKD3 to be upregulated in TNBC where it drives proliferation, cell motility and cancer stem cell survival. By mapping the PKD3 interactome and kinome our goal is to identify druggable signaling nodes for this cancer type.
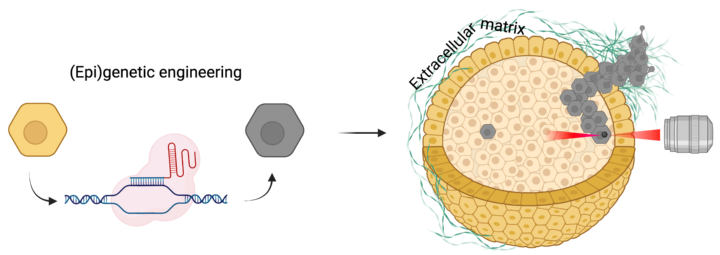
Loss of tumor suppressors and activation of oncogenes are key events in cancer development. While these events can spontaneously occur in multiple cells, only a subset of these cells will eventually contribute to tumor formation or promote metastasis. The conditions that facilitate this, particularly in 3D environments in which cells typically grow within tissues, are poorly understood. This interdisciplinary project tackles this topic by combining state of the art CRISPR/Cas9 epigenetic editing, cancer cell biology and biophysical methodologies. This project is performed in close collaboration with the Guo lab of cell mechanics (MIT) and is funded by the MISTI Global Seed Fund – University of Stuttgart program.
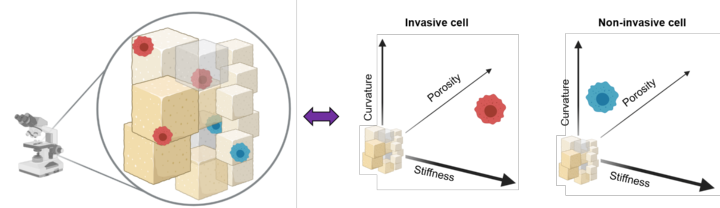
More than 70% of breast cancer associated deaths hail from the metastatic spread of the disease to distant organs. Interestingly, most breast cancer metastases affect the bones, the mechanisms involved in this tissue-specific enrichment being still poorly understood. In particular, the role of biophysical cues such as tissue topography and stiffness in this process are not clear. In this interdisciplinary project we will develop high resolution bone mimetic scaffolds to assay the metastatic colonization process. This work is performed together with the Heymann and Hörning groups from the IBBS. The initiative is funded by the Research Seed Capital Program (MWK Baden-Württemberg & University of Stuttgart).
Our Partners
We work in close collaboration with Prof. Matthias Schwab (Margarete Fischer Bosch Institute of Clinical Pharmacology, Stuttgart), Prof. Walter Aulitzky (Robert Bosch Hospital, Stuttgart), Prof. Christine Sers (Charité, Berlin), Prof. Tilman Brummer and Prof. Melanie Boerries (University of Freiburg), Prof. Hauke Busch (University of Luebeck), and Prof. Boris Macek (Proteome Center Tübingen).
Publications
Click here for a full publication list.
News
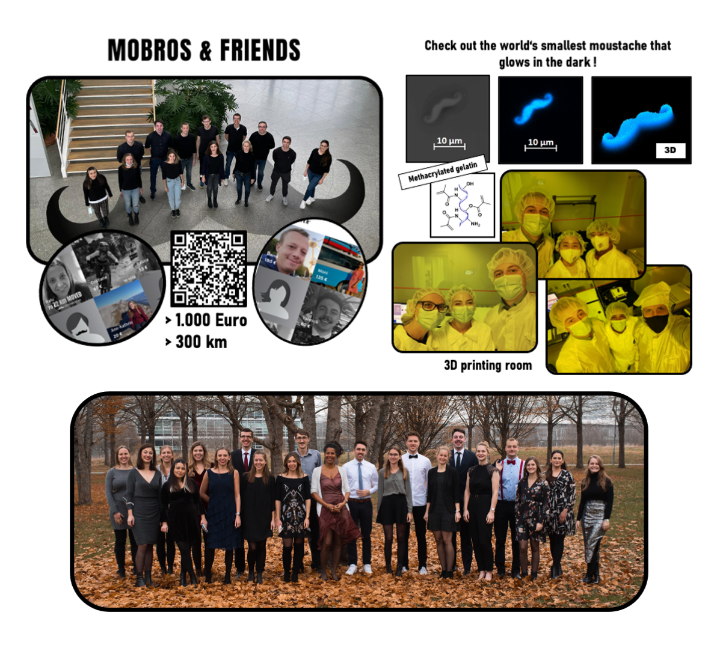
Members of the Monilola Olayioye group, or MoLab in short, shared their interest in cancer research beyond the lab, by gathering forces with friends and colleagues from the Institute of Cell Biology and Immunology (IZI) and The Institute of Biomaterials and Biomolecular Systems (IBBS) to join Movember 2021. The MoBros & Friends strong and determined team set out to promote the mission of improving men’s health worldwide, which includes research on prostate cancer, through fun and healthy activities. Moustaches were grown, more than 300 km walked, healthy snacks were baked, and last but not least, the world’s smallest “glow-in-the-dark” moustache was printed thanks to our friends and 3R-US collaborators at IBBS, especially PhD student Kai Hirzel in the group of Michael Heymann. All team progress was monitored and shared with the world, and as a result, more than 1000€ was raised for the cause. Congratulations to the team, and we hope to top the success next year!
Monilola Olayioye (Lab Head)
Cristiana Lungu (Postdoctoral Scientist, RiSC and MISTI PI)
Bettina Noll (Postdoctoral Scientist)
Lisa Brenner (PhD student)
Merih Özverin (PhD student)
Florian Meyer (PhD student)
Stella Asmanidou (joint PhD student with the Kontermann lab)
Fiona Kühnel (PhD student)
Ann-Kathrin Löffler (joint PhD student with the Kontermann lab)
Nina Pelzer (Master student)
Lisa Tellier (exchange MSc student Montpellier)
Nikolai Herdrich (student assistant)
Lisa Johanna Lang (student assistant)
Simone Schmid (Technician)
How to join our Group
Undergraduate students interested in conducting a BSc/MSc research project should get in touch at an early stage with their CV as space is limited. PhD and postdoctoral positions are advertised on our career portal JoinUS. We are happy to support highly qualified and competitive candidates that wish to apply for fellowship funding programs to join our research team.
Stellenbeschreibung
In der Abteilung Molekulare Tumorzellbiologie am Institut für Zellbiologie und Immunologie der Universität Stuttgart (https://www.izi.uni-stuttgart.de/en/research/olayioye/) ist zum nächstmöglichen Zeitpunkt die unbefristete Stelle einer Biologisch-Technischen Laborassistenz (w/m/d) mit einem Stellenumfang von 50% zu besetzen. Die Vergütung erfolgt nach TV-L bis Entgeltgruppe 9, je nach Vorliegen der persönlichen und tarifrechtlichen Voraussetzungen. Die Arbeitszeiten können flexibel vereinbart werden. Die Stelle soll das laufende 3R-US Projekt unterstützen.
Ihre Aufgabengebiete
- Mitwirkung im Bestellwesen, Lagerverwaltung/Vorratshaltung
- Aufrechterhaltung der allgemeinen Labororganisation und Instandhaltung von Laborgeräten
- Bereitstellen von Medien und Reagenzien
- Pflege der Biobank: Anlegen, Halten und Charakterisierung von Stammzellkulturen aus humanem Material und aus der Maus
- Arbeiten mit Zellkulturen und Geweben (immunhistologische und mikroskopische Untersuchungen)
- Durchführung von molekular- und zellbiologischen Assays
- Proteinchemische Arbeiten
- Etablierung von Praktikumsversuchen und Durchführung von studentischen Praktika
- Labortechnische Mitbetreuung von Studierenden im Rahmen von BSc/MSc Abschlussarbeiten
Ihr optimales Profil
- eine abgeschlossene Ausbildung als biologisch-technische Assistenz oder eine vergleichbare Qualifikation
- praktische Erfahrung in der Kultivierung von Zellen und Geweben und deren Charakterisierung durch histologische Verfahren
- Erfahrung in genetischen, molekularbiologischen und biochemischen Arbeitsmethoden sind von Vorteil (z.B. PCR, DNA, RNA und Protein Isolation)
- EDV-Kenntnisse (Labor-EDV, Excel, MS-Word, Powerpoint)
- Bereitschaft für das Erlernen neuer Untersuchungsmethoden
- Zuverlässigkeit, Flexibilität, Team- und Kommunikationsfähigkeit (idealerweise auch in englischer Sprache
Wir halten einen modernen und abwechslungsreichen Arbeitsplatz in einem engagierten, internationalen Team für Sie bereit. Möglichkeiten zur Weiterbildung werden Ihnen über das ZLW der Universität Stuttgart angeboten. Bitte senden Sie Ihre aussagekräftige Bewerbung per PDF bis zum 22.10.2023 an: monilola.olayioye@izi.uni-stuttgart. Für Rückfragen steht Ihnen Frau Gabi Sawall im Institutssekretariat gerne zur Verfügung (0711 685 66988).
Die Universität Stuttgart legt Wert auf die berufliche Gleichstellung, deshalb werden schwerbehinderte Bewerber*innen bei gleicher Eignung vorrangig berücksichtigt. Die Bewerbung von Frauen ist ausdrücklich erwünscht. Die Einstellung erfolgt durch die Zentrale Verwaltung der Universität. Informationen zur Erhebung von personenbezogenen Daten nach Artikel 13DS-GVO finden Sie unter https://www.uni-stuttgart.de/datenschutz/bewerbung. Bitte beachten Sie bei der Übersendung Ihrer Bewerbung per E-Mail, dass bei diesem Übermittlungsweg Ihre Daten unverschlüsselt sind und unter Umständen von Unbefugten zur Kenntnis genommen oder auch verfälscht werden können. Gerne können Sie uns Ihre Unterlagen per Post zukommen lassen. Bewerbungs- und Vorstellungskosten können leider nicht erstattet werden.
Wir freuen uns auf Ihre Bewerbung!
Download: TA Position 3R Uni Stuttgart